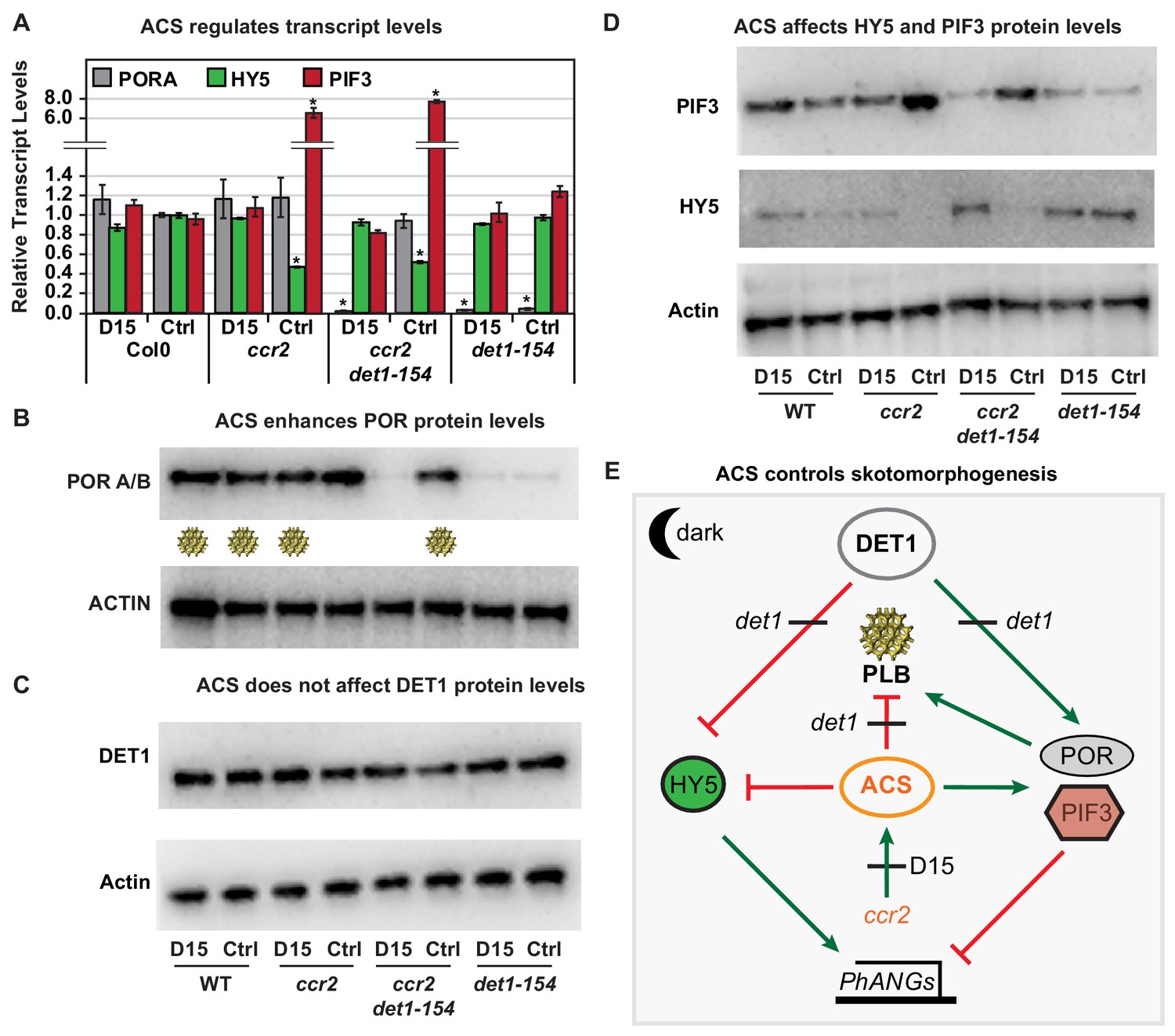
Retrograde Control of Cytosolic Translation Targets Synthesis of Plastid Proteins and Nuclear Responses for High-Light Acclimation | bioRxiv

Phytochrome and retrograde signalling pathways converge to antagonistically regulate a light-induced transcriptional network | Nature Communications

Toward an Integrated Understanding of Retrograde Control of Photosynthesis | Antioxidants & Redox Signaling
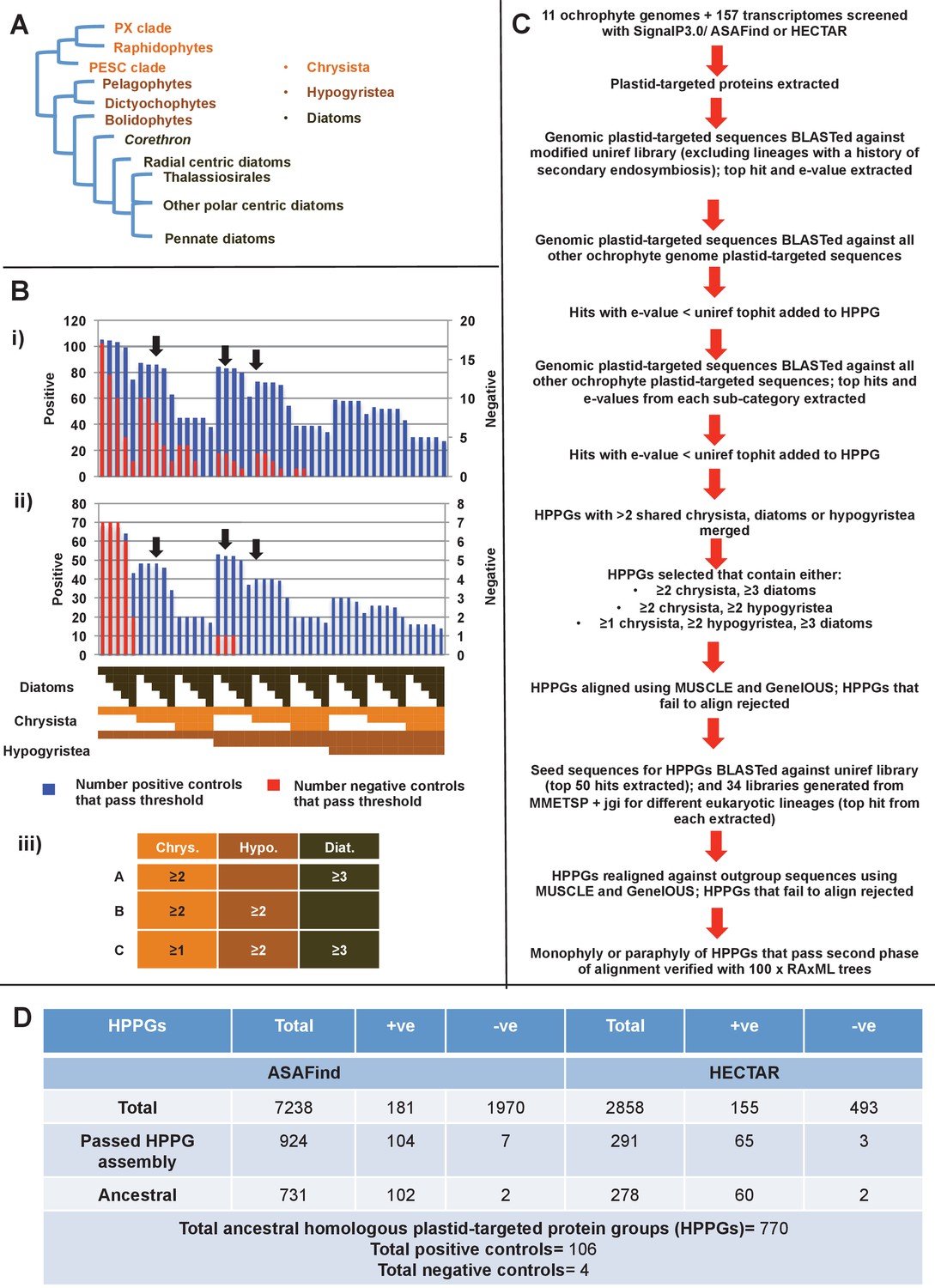
Chimeric origins of ochrophytes and haptophytes revealed through an ancient plastid proteome | eLife

Evolutionary analysis of Arabidopsis, cyanobacterial, and chloroplast genomes reveals plastid phylogeny and thousands of cyanobacterial genes in the nucleus | PNAS

Retrograde Control of Cytosolic Translation Targets Synthesis of Plastid Proteins and Nuclear Responses for High-Light Acclimation | bioRxiv

Plant thiol peroxidases as redox sensors and signal transducers in abiotic stress acclimation - ScienceDirect

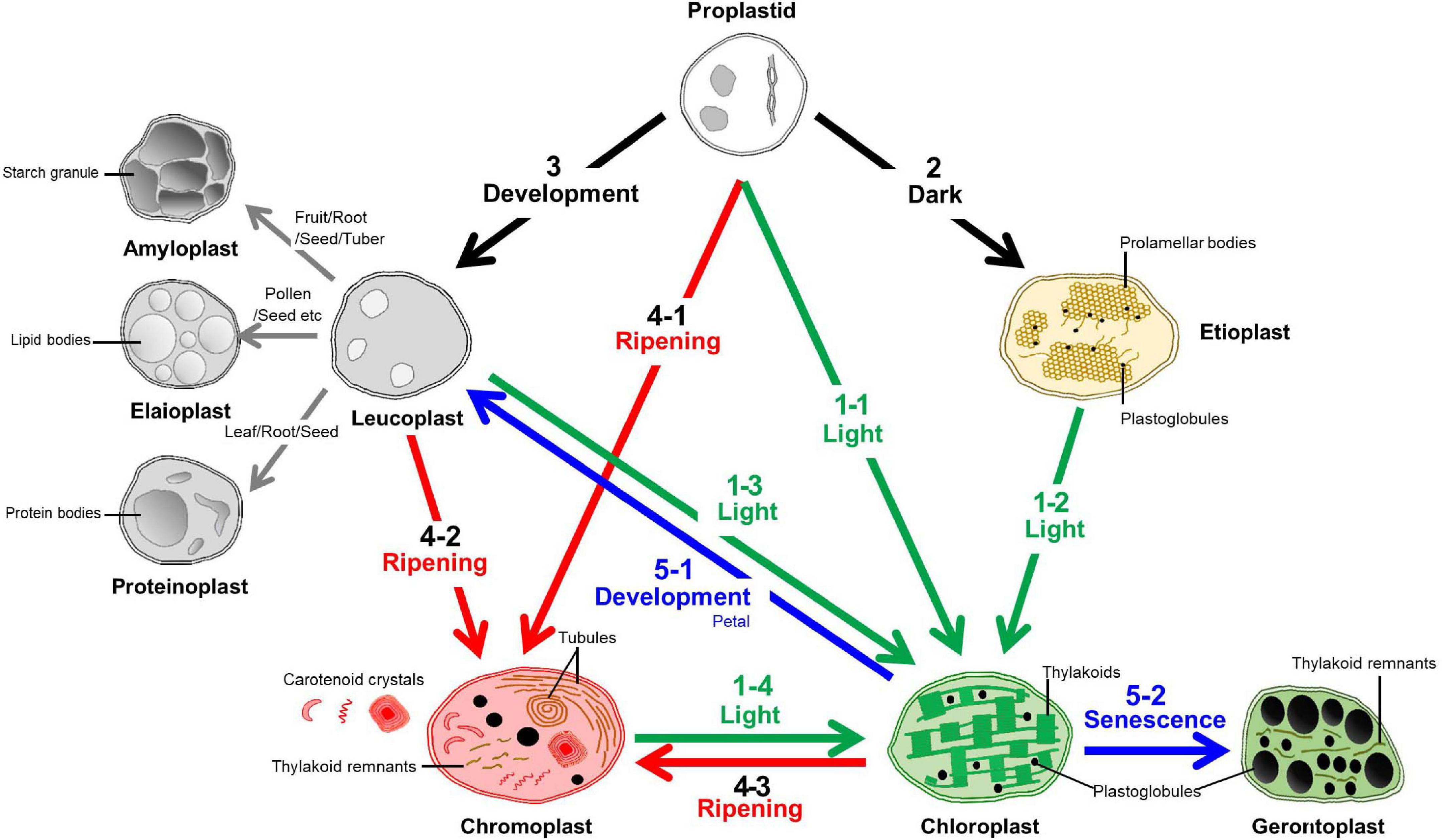
The impact of ABA on plastid differentiation in 7-day-old light-grown... | Download Scientific Diagram
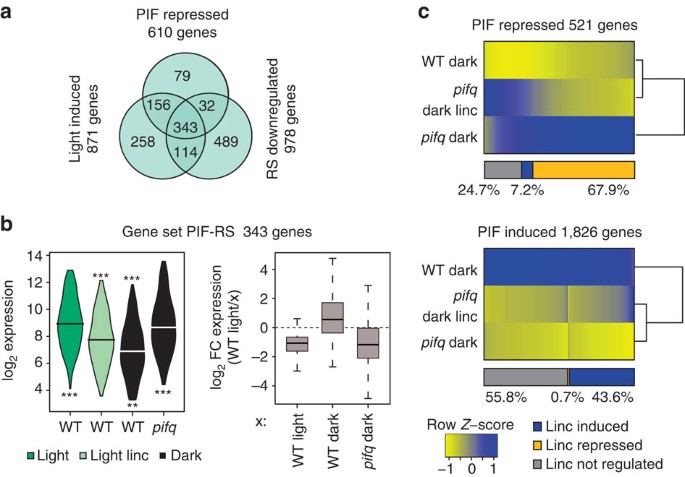
Phytochrome and retrograde signalling pathways converge to antagonistically regulate a light-induced transcriptional network | Nature Communications
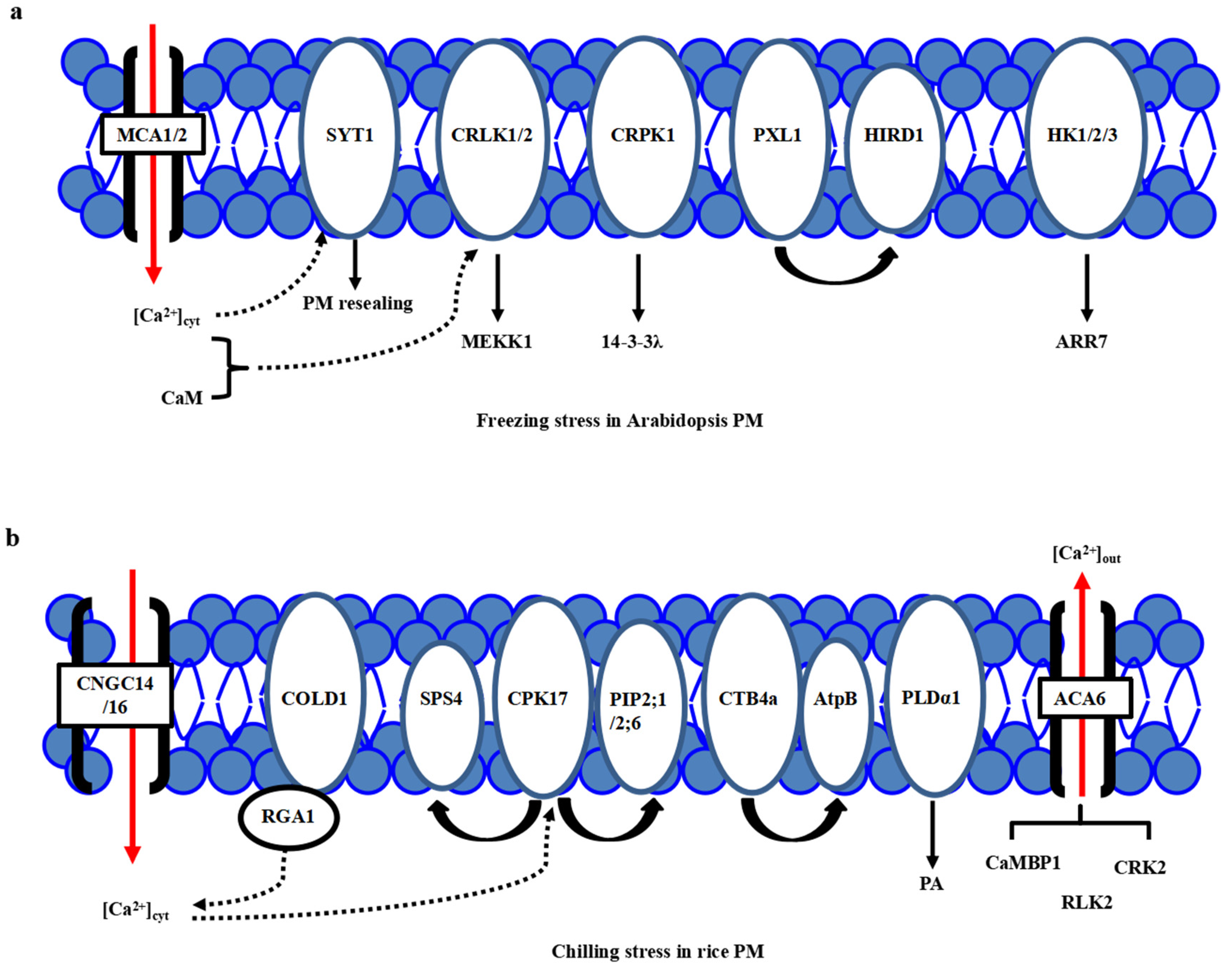
Plants | Free Full-Text | Convergence and Divergence: Signal Perception and Transduction Mechanisms of Cold Stress in Arabidopsis and Rice
Vesicles Bearing Toxoplasma Apicoplast Membrane Proteins Persist Following Loss of the Relict Plastid or Golgi Body Disruption | PLOS ONE
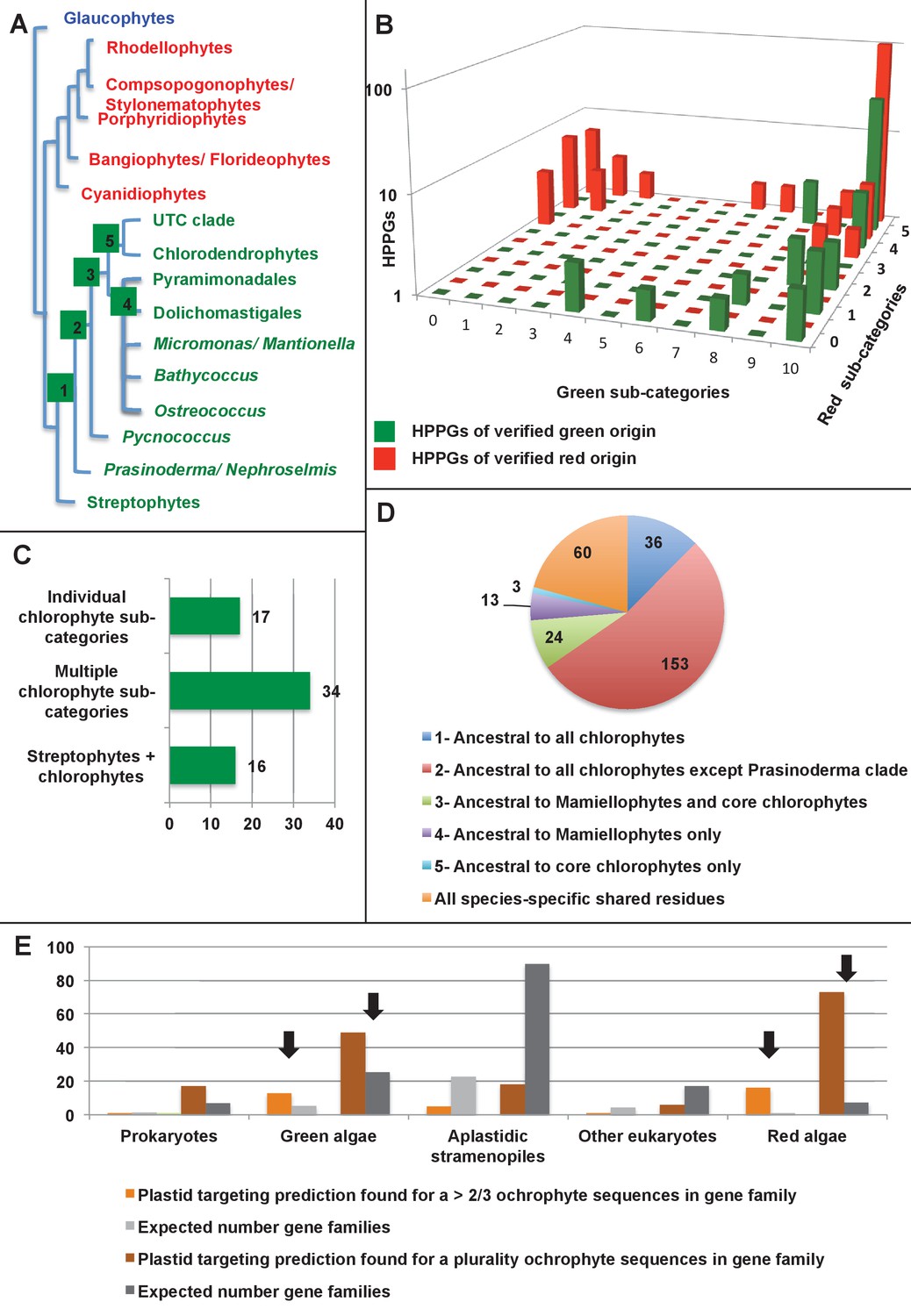
Chimeric origins of ochrophytes and haptophytes revealed through an ancient plastid proteome | eLife

PDF) Genome-wide signatures of plastid-nuclear coevolution point to repeated perturbations of plastid proteostasis systems across angiosperms